code-cells 
- Description
- Lightweight notebooks with support for ipynb files
- Latest
- code-cells-0.5.tar (.sig), 2024-Nov-19, 60.0 KiB
- Maintainer
- Augusto Stoffel <arstoffel@gmail.com>
- Website
- https://github.com/astoff/code-cells.el
- Browse ELPA's repository
- CGit or Gitweb
- Badge
To install this package from Emacs, use package-install or list-packages.
Full description
This package lets you efficiently navigate, edit and execute code split into cells according to certain magic comments. Moreover, if you have Jupytext or Pandoc installed, you can also open ipynb notebook files directly in Emacs. They will be automatically converted to a script for editing, and converted back to notebook format when saving.
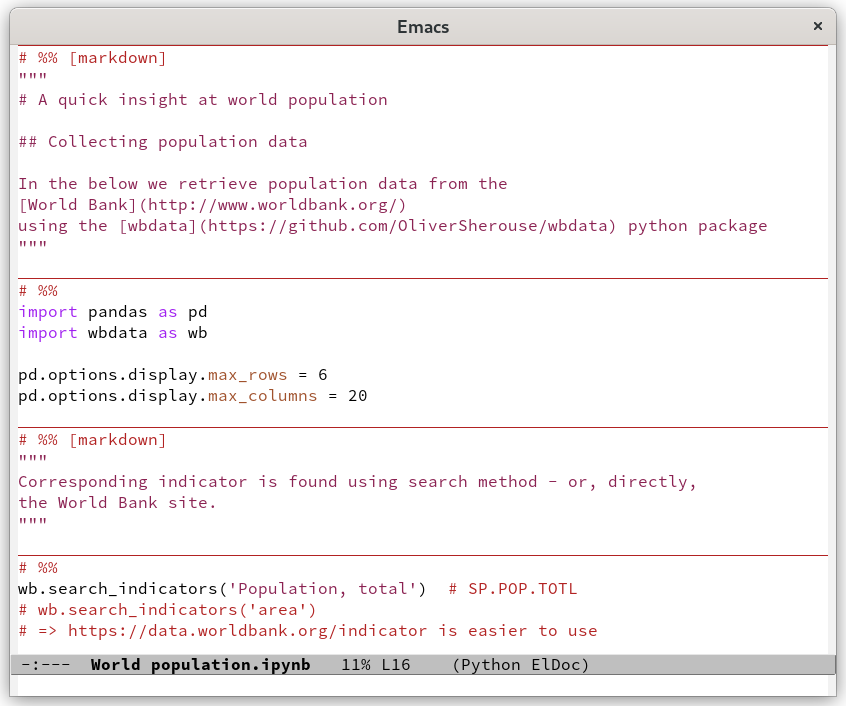
Figure 1: Working on a Jupyter notebook as a plain Python script
By default, the following kinds of comment lines are recognized as cell boundaries:
# In[<number>]: # %% Optional title
The first is what you get by exporting a notebook to a script on
Jupyter's web interface or with the command jupyter nbconvert. The
second style is compatible with Jupytext, among several other tools.
More percent signs signify nested cells. See section “Customization”
below to learn how to change the cell boundary pattern.
Advertisement: To complement this package, you may be interested in dREPL, a fully featured shell for Python and other languages with graphical capabilities.
1. Minor mode
The code-cells-mode minor mode provides the following things:
- Fontification of cell boundaries.
- Keybindings for the cell navigation and evaluation commands, under the
C-c %prefix. - Outline mode integration: cell headers have outline level determined
by the number of percent signs or asterisks; within a cell, outline
headings are as determined by the major mode, but they are demoted
by an amount corresponding to the level of the containing cell.
This provides code folding and hierarchical navigation, among other
things, when
outline-minor-modeis active.
code-cells-mode is automatically activated when opening an ipynb
file, but of course you can activate it in any other buffer, either
manually or through some hook. There is also the
code-cells-mode-maybe function, which activates the minor mode if
the current buffer seems to contain cell boundaries. It can be used
like this, for instance:
(add-hook 'python-mode-hook 'code-cells-mode-maybe)
2. Editing commands
This package provides a number of cell editing commands. They are
listed below together with their bindings in code-cells-mode-map.
Note, however, that these commands do not require the minor mode to be
active; you can bind them to other keys or call them via M-x
anywhere you like.
code-cells-backward-cell(C-c % p)code-cells-forward-cell(C-c % n)code-cells-move-cell-up(C-c % P)code-cells-move-cell-down(C-c % N)code-cells-comment-or-uncomment(C-c % ;)code-cells-copy(C-c % w)code-cells-deletecode-cells-duplicate(C-c % d)code-cells-eval(C-c % e)code-cells-eval-and-step(C-c % s)code-cells-eval-above(C-c % a)code-cells-eval-belowcode-cells-eval-whole-buffercode-cells-indent(C-c % \)code-cells-kill(C-c % C-w)code-cells-mark-cell(C-c % @)
The code-cells-eval command sends the current cell to a suitable
REPL, chosen according to the current major and minor modes. The
exact behavior is controlled by the code-cells-eval-region-commands
variable, which can be customized to suit your needs.
You may prefer shorter keybindings for some of these commands. One
sensible possibility is to use C-c C-c to evaluate and M-p
resp. M-n to navigate cells. This can be achieved with the
following configuration:
(with-eval-after-load 'code-cells
(let ((map code-cells-mode-map))
(keymap-set map "M-p" 'code-cells-backward-cell)
(keymap-set map "M-n" 'code-cells-forward-cell)
(keymap-set map "C-c C-c" 'code-cells-eval)
;; Overriding other minor mode bindings requires some insistence...
(keymap-set map "<remap> <jupyter-eval-line-or-region>" 'code-cells-eval)))
Finally, a remark on prefix arguments. For most commands, the logic is that the absolute value of the numeric prefix stipulates the number of cells around the current cell to act on, with the sign determining whether to include cells above or below it. The numeric prefixes 0 and -1 have a special meaning: they tell to act on the half of the current cell below respective above the current line.
3. Customization
Type M-x customize-group RET code-cells RET to see a listing of all
customization options.
The most important option is code-cells-boundary-regexp, which
determines which lines of a buffer should be regarded as a cell
boundary. Some alternatives to the default value include:
"^#\\(#+\\)": This recognizes lines starting with 2 or more hash characters as cell boundaries, which is an interesting option, say, for Python:## Cell title (optional) # Some commentary, which normally starts with # a single hash character. print("hello")"^;;\\(;+\\) ": This recognizes lines starting with 3 or more semicolons as cell boundaries, which is an interesting option, say, for Emacs Lisp code split into sections:;;; Section title ;; Some commentary, which by convention starts ;; with double semicolons. (message "hello")
"^\\s<+\\(\\*+\\)": This regular expression recognizes lines of the following form as cell boundaries:#* #** #***
This implements a kind of "reverse literate programming" where the prose part is behind comments and can have Org-like syntax (the number of asterisks determines the heading level).
As usual, you can customize code-cells-boundary-regexp globally, or
change it for a single major mode, for instance with
(add-hook 'emacs-lisp-mode-hook (lambda () (setq-local code-cells-boundary-regexp "^;;\\(;+\\)")))
or even modify it in a single project using directory-local variables, e.g. by typing the following:
M-x add-dir-local-variable RET python-mode RET code-cells-boundary-regexp RET "^#\\(#+\\)" RET
Note: Until version 0.4, the third cell boundary style above was included in the default settings. Use the suggested customization to recover the old behavior.
4. Speed keys
Similarly to Org mode's speed keys, the code-cells-speed-key
function returns a key definition that only acts when the point is at
the beginning of a cell boundary. Since this is usually not an
interesting place to insert text, you can assign short keybindings
there.
No speed keys are set up by default. A sample configuration is as follows:
(with-eval-after-load 'code-cells
(let ((map code-cells-mode-map))
(define-key map "n" (code-cells-speed-key 'code-cells-forward-cell))
(define-key map "p" (code-cells-speed-key 'code-cells-backward-cell))
(define-key map "e" (code-cells-speed-key 'code-cells-eval))
(define-key map (kbd "TAB") (code-cells-speed-key 'outline-cycle))))
For Evil users, the following can be used:
(with-eval-after-load 'code-cells
(let ((map code-cells-mode-map))
(define-key map [remap evil-search-next] (code-cells-speed-key 'code-cells-forward-cell)) ;; n
(define-key map [remap evil-paste-after] (code-cells-speed-key 'code-cells-backward-cell)) ;; p
(define-key map [remap evil-backward-word-begin] (code-cells-speed-key 'code-cells-eval-above)) ;; b
(define-key map [remap evil-forward-word-end] (code-cells-speed-key 'code-cells-eval)) ;; e
(define-key map [remap evil-jump-forward] (code-cells-speed-key 'outline-cycle)))) ;; TAB
5. Handling Jupyter notebook files
With this package, you can edit Jupyter notebook (*.ipynb) files as
if they were normal plain-text scripts. Converting to and from the
JSON-based ipynb format is done by an external tool, Jupytext by
default, which needs to be installed separately.
Note that the result cells of ipynb files are not retained in the conversion to script format. This means that opening and then saving an ipynb file clears all cell outputs.
While editing a converted ipynb buffer, you can use the regular
write-file command (C-x C-w) to save a copy in script format, as
displayed on the screen. Moreover, from any script file with cell
separators understood by Jupytext, you can call
code-cells-write-ipynb to save a copy in notebook format.
5.1. Tweaking the ipynb conversion
If relegating markdown cells to comment blocks offends your literate programmer sensibilities, try including the following in the YAML header of a converted notebook (and then save and revert it). It will cause text cells to be displayed as multiline comments.
jupyter:
jupytext:
cell_markers: '"""'
It is also possible to convert notebooks to markdown or Org mode format. For markdown, use the following:
(setq code-cells-convert-ipynb-style '(("jupytext" "--to" "ipynb" "--from" "markdown")
("jupytext" "--to" "markdown" "--from" "ipynb")
(lambda () #'markdown-mode)))
To edit ipynb files as Org documents, try using Pandoc with the configuration below. In combination with org-babel, this can provide a more notebook-like experience, with interspersed code and results.
(setq code-cells-convert-ipynb-style '(("pandoc" "--to" "ipynb" "--from" "org")
("pandoc" "--to" "org" "--from" "ipynb")
(lambda () #'org-mode)))
A good reason to stick with Jupytext, though, is that it offers round-trip consistency: if you save a script and then revert the buffer, the buffer shouldn't change. With other tools, you may get some surprises.
6. Alternatives
python-cell.el provides similar cell editing commands. It seems to be limited to Python code.
With Jupytext's paired notebook mode it is possible to keep a notebook open in JupyterLab and simultaneously edit a script version in an external text editor.
The EIN package allows to open ipynb files directly in Emacs with an
UI similar to Jupyter notebooks. Note that EIN also registers major
modes for ipynb files; when installing both packages at the same time,
you may need to adjust your auto-mode-alist manually.
7. Contributing
Discussions, suggestions and code contributions are welcome! Since this package is part of GNU ELPA, nontrivial contributions (above 15 lines of code) require a copyright assignment to the FSF.
Old versions
| code-cells-0.4.tar.lz | 2024-Mar-31 | 8.79 KiB |
| code-cells-0.3.tar.lz | 2022-Sep-17 | 8.64 KiB |
| code-cells-0.2.tar.lz | 2022-Feb-28 | 7.38 KiB |
News
Version 0.5 - Several new editing commands. - Integrate with repeat-mode and context-menu-mode. - Some changed keybindings. - More consistent handling of numeric arguments and cell ranges.